Dynamic PFB Importer
Overview
Dynamic PFB Importer is a new end-to-end import experience that moves data from Gen3 BioData Catalyst into BioData Catalyst powered by Seven Bridges and automatically generates a complete set of reusable project outputs. Unlike earlier importer flows that primarily transferred the PFB (Avro) file itself, Dynamic PFB Importer also resolves DRS references and downloads the linked files into the destination project. This gives users both the structured PFB metadata and the underlying file data in a single workflow.
The importer is called "dynamic" because it expands the imported PFB file into multiple immediately usable outputs, including schema files, flattened metadata files, node-specific TSVs, a DRS manifest, and downloaded files arranged in preserved folder paths when path information is available.
Learn more about PFB at the NHLBI BioData Catalyst® (BDC) Documentation site – Overview of the Portable Format for Bioinformatics (PFB) file type
What this feature provides
• A direct export path from Gen3 BioData Catalyst into BioData Catalyst powered by Seven Bridges.
• Automatic import of the original PFB Avro file into the destination project.
• Automatic generation of schema and metadata outputs derived from the imported Avro.
• Automatic creation of one TSV per node type so graph content is easier to review and reuse.
• Automatic generation of a DRS manifest that captures valid DRS-linked file entries.
• Automatic download of the data referenced by DRS URIs, not just the metadata that points to those files.
• Preservation of folder structure for downloaded DRS files when destination path information is provided by the source.
• A single import flow that gives users both project-ready metadata outputs and file-level data outputs.
Before you begin
Before starting an import, make sure the following conditions are met:
• You can sign in to Gen3 BioData Catalyst and access the Exploration page.
• You can sign in to BioData Catalyst powered by Seven Bridges and have permission to import into at least one destination project.
• Your destination project is appropriate for the data being imported, especially if the import includes controlled-access data.
• You have sufficient storage and permissions to view imported files, folders, metadata, and tags after the import completes.
How the Dynamic PFB Importer works
The workflow begins in Gen3 BioData Catalyst and finishes in BioData Catalyst powered by Seven Bridges. Users select the data to export, choose BioData Catalyst powered by Seven Bridges as the destination, and complete the import in BioData Catalyst powered by Seven Bridges by selecting a project and optionally adding tags. Once the import starts, the platform performs the downstream processing automatically.
1. Select data in Gen3 BioData Catalyst
- Sign in to Gen3 BioData Catalyst: NHLBI BioData Catalyst®

GEN3 BioData Catalyst homepage
- Open the Exploration page.
Exploration tab
- Browse or filter the available studies and files.
- Select the data items you want to export.
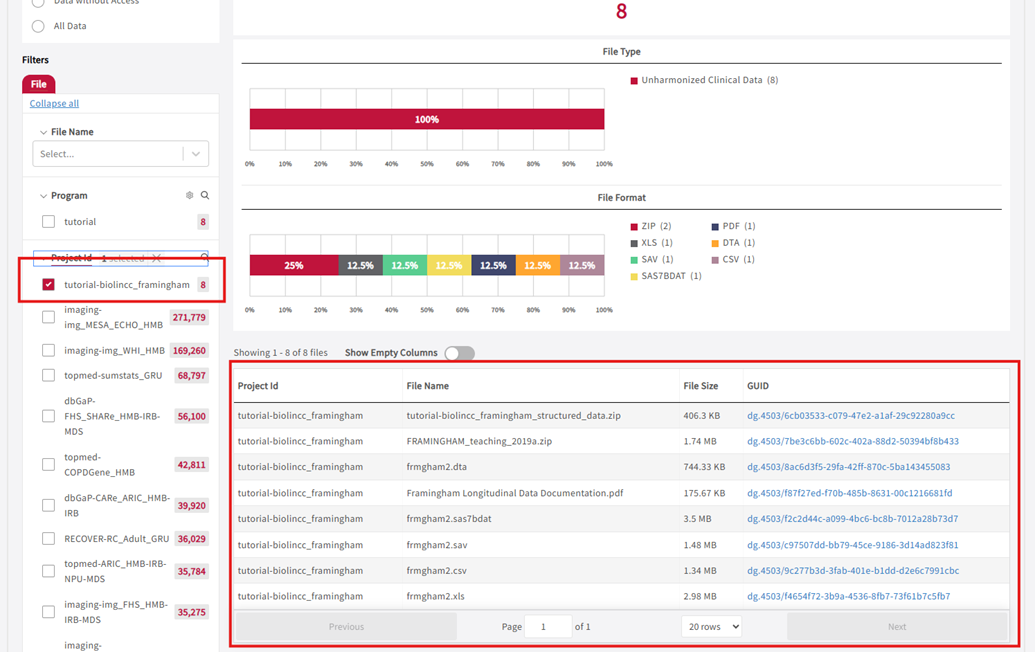
Project ID: tutorial-biolincc_framingham with 8 files selected
- Open Export to Seven Bridges and choose Export to BioData Catalyst powered by Seven Bridges.
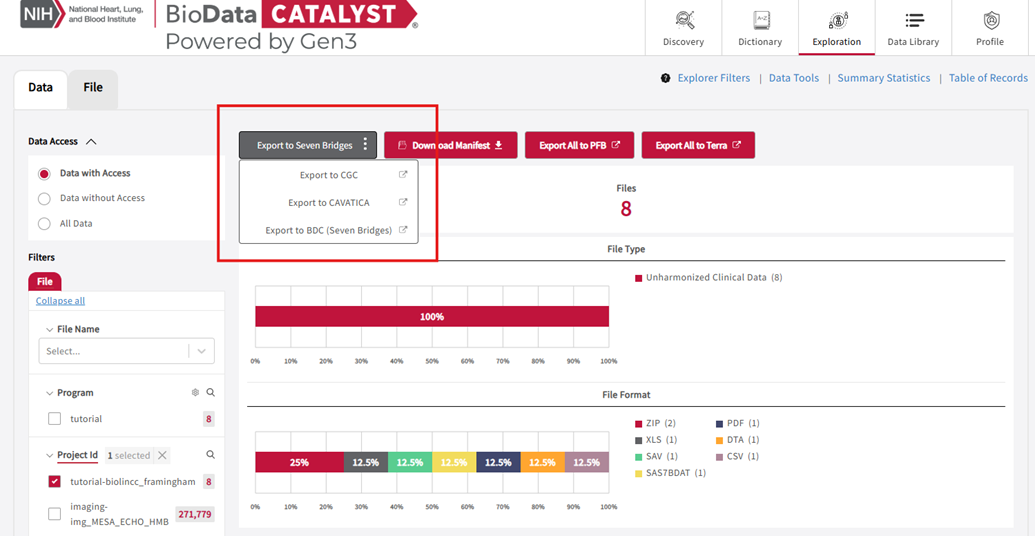
Export to Seven Bridges options
After you choose Export to BioData Catalyst powered by Seven Bridges, the platform redirects you into the platform import flow. The selected source context is carried forward so the user can continue the import without rebuilding the selection manually.
2. Complete the import in BioData Catalyst powered by Seven Bridges
- Sign in to BioData Catalyst powered by Seven Bridges if prompted.
It may prompt you to link your BDC account to GEN3
- Review the import context preloaded from Gen3 BioData Catalyst.
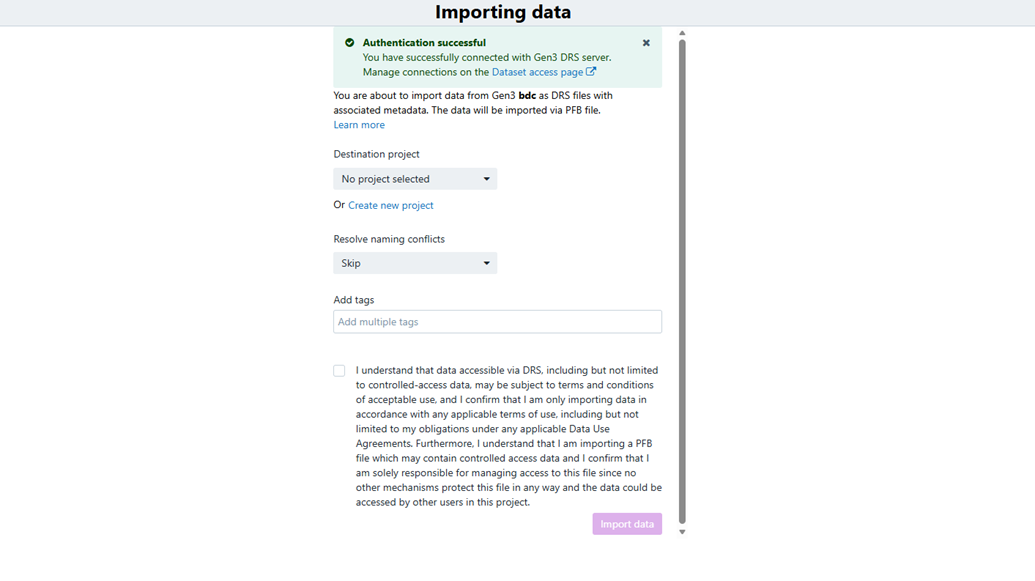
Importing data
- Choose an existing destination project or create a new one.
- Add tags that should be applied to the imported content.
- Click Import Data to start the importer.
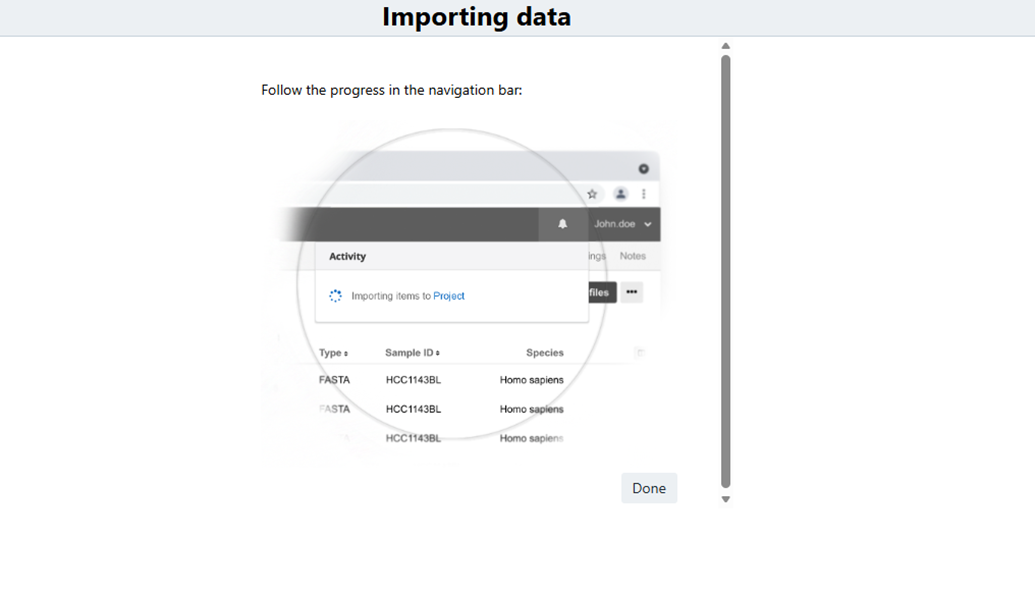
Importing data
When the import begins, Dynamic PFB Importer stores the original PFB input and generates the additional output artifacts automatically. Users do not need to run separate post-processing steps to obtain the derived metadata files, node-type TSVs, or DRS-linked file downloads.
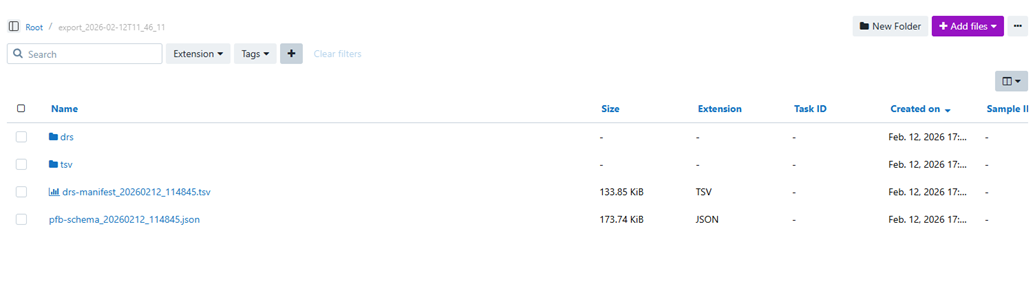
What appears in the destination project
After the import completes, the destination project contains the original input manifest together with a structured set of generated outputs. File names may include a timestamp token in the form <datetime>. This is expected and helps distinguish one import run from another.
/project-folder/
├── input-file.avro
└── output-folder/
├── pfb-schema<datetime>.json
├── drs-manifest<date_time>.tsv
├── drs/
│ ├── drs-file-1
│ ├── drs-file-2
│ └── ...
└── tsv/
├── metadata.tsv
├── node-type-A.tsv
├── node-type-B.tsv
└── ...
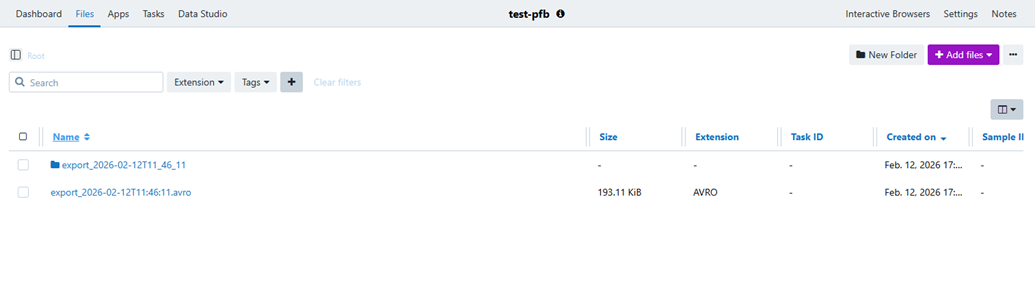
Generated outputs explained
| Output | What it contains | Why it is useful |
|---|---|---|
| input-file.avro | The original imported PFB manifest in Avro format. | Serves as the source input for the entire import process and preserves the exact payload used to generate all downstream outputs. |
| pfb-schema_<date_time>.json | A JSON representation of the Avro schema, including records, arrays, enums, and field definitions. | Provides a human readable view of the PFB structure, useful for validation, debugging, and documentation without needing Avro tools. |
| drs-manifest_<date_time>.tsv | A tab delimited manifest listing all valid DRS URIs extracted from the input, along with associated metadata fields. | Enables auditing of which DRS entries were processed and supports reuse as an import manifest in downstream file operations. |
| tsv/metadata.tsv | A flattened, tab delimited version of metadata extracted from the PFB (names, values, properties, links, ontology references where applicable). | Offers a spreadsheet friendly format for review, filtering, QC checks, and analysis. Flattened structure ensures easy consumption in tools like Excel or scripting workflows. |
| tsv/node-type-*.tsv | One TSV per node type present in the PFB (e.g., Participant, Sample, File). Each includes attribute columns and DRS URI references where applicable. | Makes graph structured PFB data easier to inspect by splitting it into entity specific tables while preserving all per node attributes. |
| drs/ | A folder containing successfully resolved files downloaded using valid DRS URIs. | Provides the actual file content referenced in the PFB so users can directly work with the underlying data, not just metadata. |
Preserved folder structure for downloaded DRS files
One of the most important improvements in Dynamic PFB Importer is that it preserves folder paths for downloaded DRS files whenever the source provides a destination path through access_method.access_url. If a path is present, the importer recreates the matching subfolder path under drs/ and places the file in that location. If no destination path is available, the file is placed at the root of the drs/ folder.
This behavior makes imported projects easier to browse and reduces the manual work required to reorganize files after import. For users working with larger or more structured datasets, preserved paths make the imported project feel much closer to the organization of the source data.
Understanding counts and imported scope
Dynamic PFB Importer produces multiple artifact types from one source export, so counts can represent different parts of the imported scope. For example, the source selection in Gen3 BioData Catalyst may represent a cohort, set, or graph of linked entities, while the generated DRS manifest reflects the subset of records that contain valid DRS URIs, and the node-type TSVs reflect graph content grouped by entity type.
• Use the original source selection as the starting point for understanding the requested import scope.
• Use the Avro and generated metadata outputs to understand the imported PFB structure and content.
• Use the DRS manifest to understand which valid DRS-linked entries were available for file download.
• Use the drs/ folder to see the file objects that were actually downloaded into the project.
• Use the tsv/ folder to review imported content by node type rather than by file count alone.
Expected behavior and format notes
• Generated file names may include a timestamp token in the form <date_time>.
• TSV outputs are UTF-8 encoded and tab-delimited.
• Flattened TSV outputs use empty cells for null values.
• Duplicate DRS URIs may appear in the DRS manifest if duplicates are present in the source content.
• Rows without a valid drs_uri are not included in the DRS manifest and do not produce downloaded files.
• file_name may be blank for a DRS entry if the source does not provide a value.
• Node-type TSVs are generated per node type and may not map one-to-one with source file counts.
Troubleshooting
Export to BioData Catalyst powered by Seven Bridges is not visible
Confirm the export is being initiated from the correct source platform and that the current environment exposes BioData Catalyst powered by Seven Bridges as an enabled destination. If platform is still not listed, contact the support team to confirm that the platform integration is enabled and that the requesting account has access to it.
The import screen does not show the Gen3 selection
Confirm that the redirect from Gen3 BioData Catalyst completed successfully and that the linked account context is available in BioData Catalyst powered by Seven Bridges.
Generated outputs do not appear in the expected project
Confirm that the import was started in the intended destination project and refresh the Files view if the interface has not yet updated.
The DRS manifest contains fewer entries than expected
The manifest includes only rows with valid DRS URIs. Records without valid DRS references are omitted from that output.
Downloaded files are not nested in subfolders
Folder preservation depends on path information being available from the source. If no destination path is provided, files appear directly under drs/.
Information to include when contacting support
• The source platform and approximate time of export.
• The destination BioData Catalyst powered by Seven Bridges project name.
• Whether the issue happened during export, redirect, import setup, or after the import completed.
• The names of the generated files or folders involved in the issue.
• A screenshot that shows the relevant part of the interface.
• Whether the issue involves generated metadata outputs, node-type TSVs, the DRS manifest, or downloaded DRS files.
Updated 2 months ago
